This web page was produced as an assignment for Genetics 564, an undergraduate capstone course at UW-Madison.
What is protein phylogeny?
Phylogeny is the study of the evolution of a subject. This subject can take the form of a species, protein, or molecule among others. Protein phylogeny relies upon the sequences of different species' respective proteins. This view on the evolution of a protein can lead to inferences on a proteins structure, function, and mechanism. (1) Many different methods can be used to construct phylogenetic trees, each with their own benefits and compromises. A couple are shown below.
How do you make a phylogenetic tree?
The first step towards constructing a phylogenetic tree is collecting sequence data for your protein. There are multiple options for this collection but the ones used to construct the phylogenetic trees shown later on this page were Ensemble and Homologene. These sequences were then aligned and phylogenetic trees were constructed using Clustal Omega and MEGA.
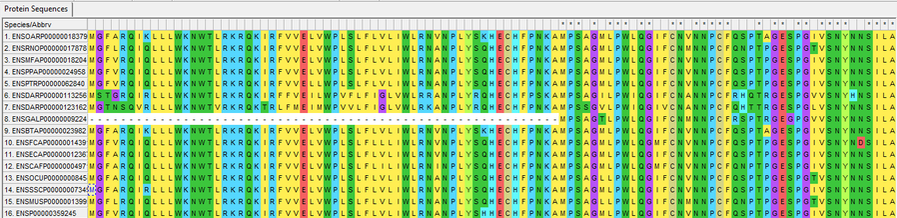
Aligned protein sequences for ABCA4.
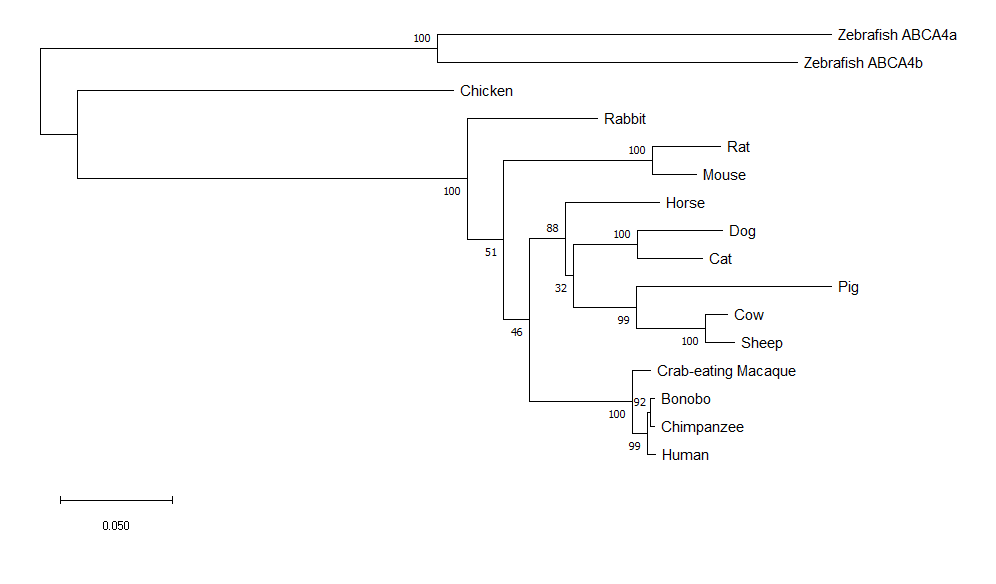
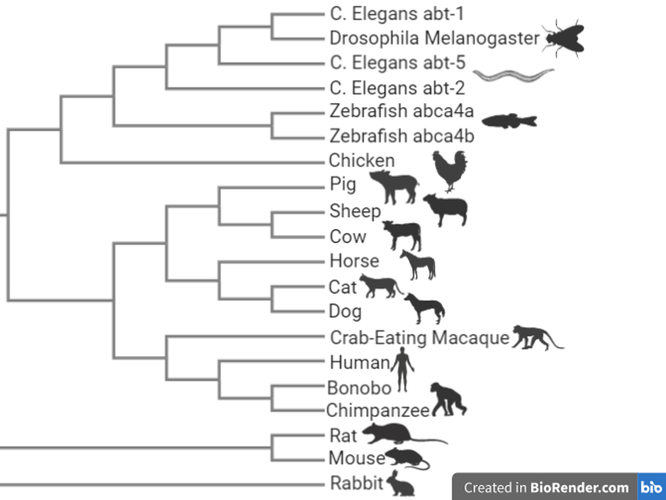
MEGA constructed maximum likelihood tree.
Maximum likelihood trees are formed through assigning probabilities to mutations instead of counts. The tree with the highest likelihood based on mutation probabilities is selected. (2)
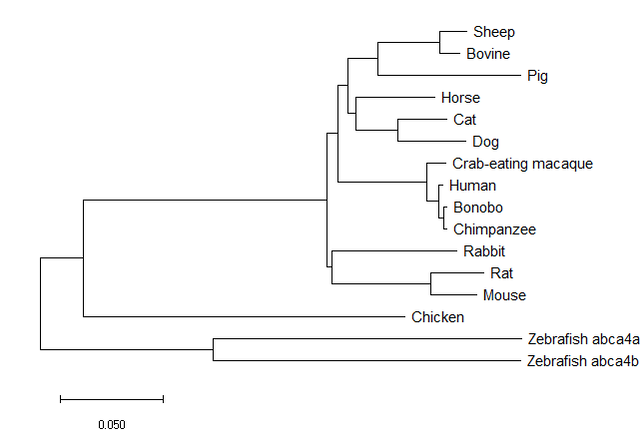
MEGA constructed neighbor-joining tree.
Neighbor-joining trees works to minimize the the branch length of each neighboring sequence, starting with a star-like tree. (3)
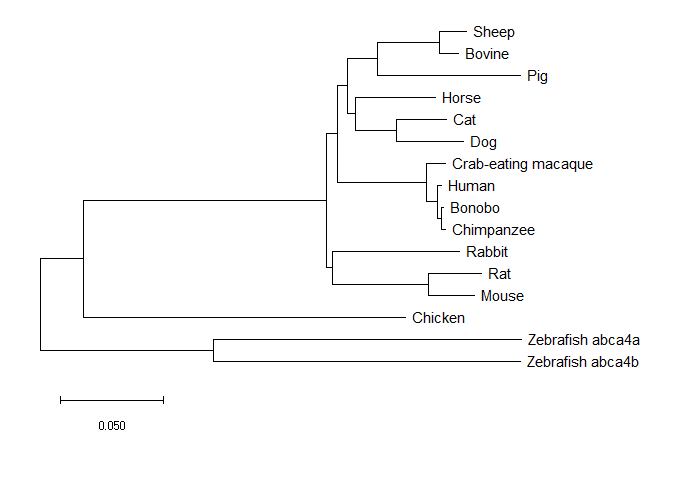
MEGA constructed minimum evolution tree.
The minimum evolution tree works by choosing shortest branch lengths from all possible branch lengths. (4)
Clustal Omega constructed tree.
Discussion
The results above show similar results, giving wide confidence in the evolutionary pathway of the ABCA4 gene. This information can be used to inform other methods, for example in cases where function is conserved despite phylogenetic distances between species.
References:
1. Chang, A. B., Lin, R., Studley, W. K., Tran, C. V., & Saier, J. M. H. (2004). Phylogeny as a guide to structure and function of membrane transport proteins (Review). Molecular Membrane Biology, 21(3), 171–181. doi: 10.1080/09687680410001720830
2. Khachatourians, G. G., & Arora, D. K. (2006). Applied mycology and biotechnology. Amsterdam: Elsevier.
3. The neighbor-joining method: a new method for reconstructing phylogenetic trees. (1987). Molecular Biology and Evolution. doi: 10.1093/oxfordjournals.molbev.a040454
4. THOMPSON, E.A. (1973), The method of minimum evolution. Annals of Human Genetics, 36: 333-340. doi:10.1111/j.1469-1809.1973.tb00595.x
2. Khachatourians, G. G., & Arora, D. K. (2006). Applied mycology and biotechnology. Amsterdam: Elsevier.
3. The neighbor-joining method: a new method for reconstructing phylogenetic trees. (1987). Molecular Biology and Evolution. doi: 10.1093/oxfordjournals.molbev.a040454
4. THOMPSON, E.A. (1973), The method of minimum evolution. Annals of Human Genetics, 36: 333-340. doi:10.1111/j.1469-1809.1973.tb00595.x