This web page was produced as an assignment for Genetics 564, an undergraduate capstone course at UW-Madison.
What is transcriptomics?
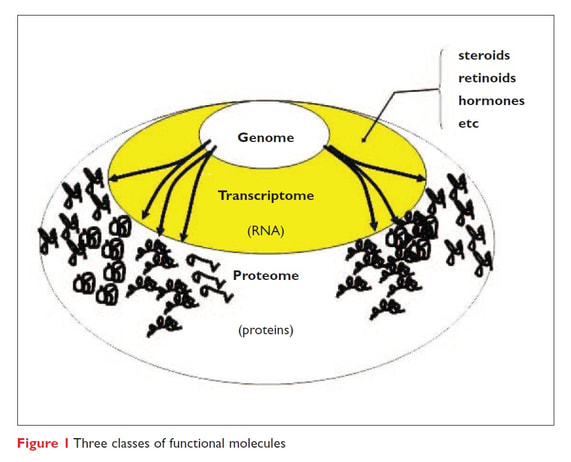
The genome of any organism produces RNA from only a portion of the DNA present in every cell (called expression), and cell function is heavily attributed to the unique expression of that particular cell. Transcriptomics is the study of RNA expression of cells, also called the transcriptome. Messenger RNA (mRNA) is the primary RNA that is analyzed in most forms of transcriptomics because mRNA is the portion of RNA produced that becomes protein through a process called translation. Protein then goes on to do most of the cells functions. Through studying the transcriptome the common production and function of a cell can be determined, alongside how the production and function of the cell changes according to certain stimulus. (1-3)
How is a transcriptome analyzed?
|
Microarray
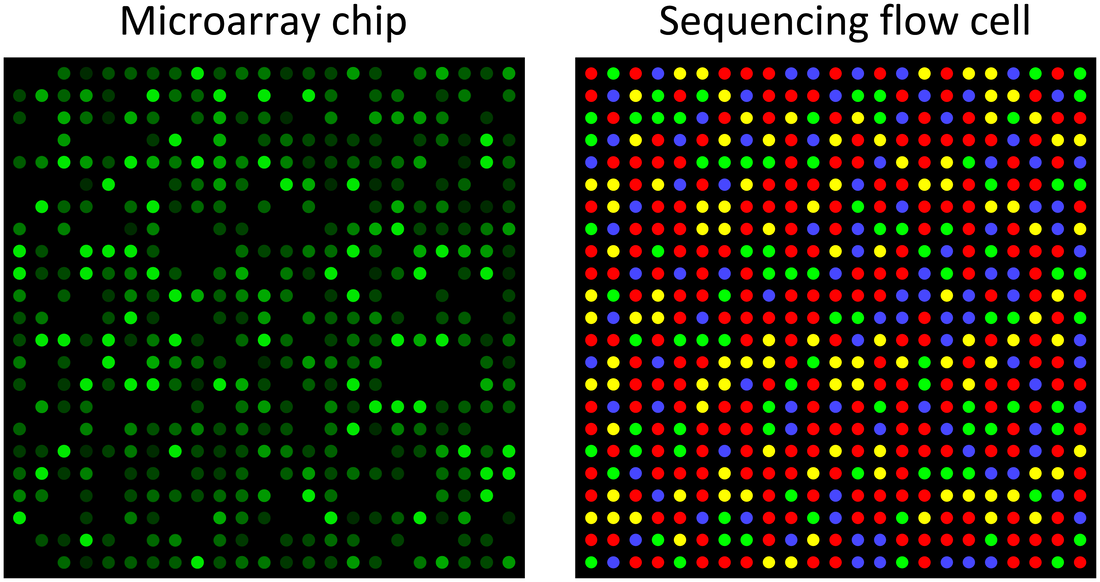
Microarrays work by having DNA sequences that researchers are interested in analyzing attached to a glass plate. DNA constructed from RNA (cDNA), with fluorescent markers attached, is washed over the slide and can be used to view expression based on intensity of fluorescence. Can not be used for discovery of new RNA sequences. (2,4)
|
RNA Sequencing
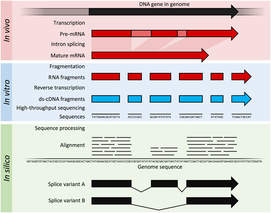
RNA sequencing works by taking a tissue of interest and isolating the RNA present. This RNA is then turned into more stable cDNA that can be used to analyze sequences. Unlike microarrays RNA sequencing can be used to discover new RNA sequences and splice variants. (2,4,5)
|
Single-Cell RNA Sequencing
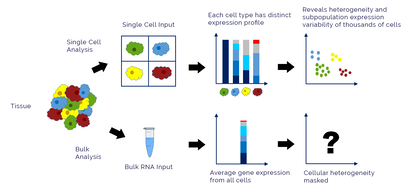
Single-Cell RNA sequencing works similarly to RNA sequencing, but allows for views into single cell type transcriptomics. This works through different methods available to researchers that isolate cells according to the interests of the researcher. It allows for further understanding an individual cells expression and function compared to RNA sequencing's average expression analysis. (5,6)
|
What can transcriptomics infer about ABCA4?
|
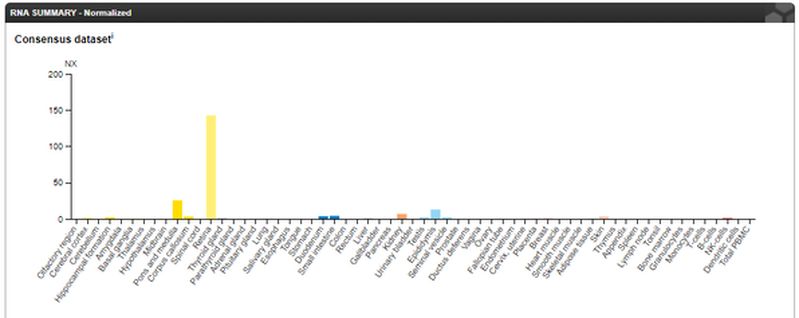
Here is data from the Human Protein Atlas, giving a summary of where ABCA4 is expressed in the body. As expected it is primarily limited to the retina, confirming it as the primary location for ABCA4 function. (7)
|
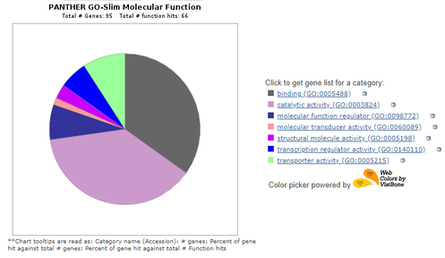
GEO Profiles, GEO DataSets, and PANTHER can be used in conjunction to analyze various aspects of genes. Here they show the various functions of ABCA4 and closely related genes operating in similar circumstances. (8-10)
|
Discussion
Presently available transcriptomics information can confirm previous assumptions and show how ABCA4's functioning has changed in response to previously recorded stimuli. Through recorded expression profiles there has been confirmation of findings about the ABCA4 gene and its functions. Transcriptomics can also be used during experimentation to analyze further how ABCA4's function changes in response to other forms of analysis and modifications that are desired in order to elucidate further the intricacies of ABCA4. This will be invaluable for further research.
References
1. Seligmann, B. (2019). TRANSCRIPTOMICS - Realising the promise with a new era of drug discovery and diagnostics. Retrieved March 19, 2020, from https://www.ddw-online.com/enabling-technologies/p148423-transcriptomics-realising-the-promise-with-a-new-era-of-drug-discovery-and-diagnostics.html
2. Blackburn, L. (2017, August 18). What is transcriptomics? Retrieved March 19, 2020, from https://www.phgfoundation.org/blog/what-is-transcriptomics
3. Transcriptomics. (n.d.). Retrieved March 19, 2020, from https://www.nature.com/subjects/transcriptomics
4. Lowe, R., Shirley, N., Bleackley, M., Dolan, S., & Shafee, T. (2017). Transcriptomics technologies. PLOS Computational Biology, 13(5). doi:10.1371/journal.pcbi.1005457
5. Single-cell RNA sequencing technologies and bioinformatics pipelines. Hwang B, Lee JH, Bang D. Exp Mol Med. 2018 Aug 7;50(8):96. doi: 10.1038/s12276-018-0071-8.
6. Single-Cell RNA Sequencing Service. (n.d.). Retrieved March 19, 2020, from https://www.lcsciences.com/discovery/applications/transcriptomics/single-cell-rna-seq-sequencing-service/
7. The Human Protein Atlas. (2020, March 06). Retrieved March 19, 2020, from https://www.proteinatlas.org/
8. Home - GEO Profiles - NCBI. (n.d.). Retrieved March 19, 2020, from https://www.ncbi.nlm.nih.gov/geoprofiles/
9. Home - GEO DataSets - NCBI. (n.d.). Retrieved March 19, 2020, from https://www.ncbi.nlm.nih.gov/gds/
10. Gene List Analysis. (n.d.). Retrieved March 19, 2020, from http://pantherdb.org/
2. Blackburn, L. (2017, August 18). What is transcriptomics? Retrieved March 19, 2020, from https://www.phgfoundation.org/blog/what-is-transcriptomics
3. Transcriptomics. (n.d.). Retrieved March 19, 2020, from https://www.nature.com/subjects/transcriptomics
4. Lowe, R., Shirley, N., Bleackley, M., Dolan, S., & Shafee, T. (2017). Transcriptomics technologies. PLOS Computational Biology, 13(5). doi:10.1371/journal.pcbi.1005457
5. Single-cell RNA sequencing technologies and bioinformatics pipelines. Hwang B, Lee JH, Bang D. Exp Mol Med. 2018 Aug 7;50(8):96. doi: 10.1038/s12276-018-0071-8.
6. Single-Cell RNA Sequencing Service. (n.d.). Retrieved March 19, 2020, from https://www.lcsciences.com/discovery/applications/transcriptomics/single-cell-rna-seq-sequencing-service/
7. The Human Protein Atlas. (2020, March 06). Retrieved March 19, 2020, from https://www.proteinatlas.org/
8. Home - GEO Profiles - NCBI. (n.d.). Retrieved March 19, 2020, from https://www.ncbi.nlm.nih.gov/geoprofiles/
9. Home - GEO DataSets - NCBI. (n.d.). Retrieved March 19, 2020, from https://www.ncbi.nlm.nih.gov/gds/
10. Gene List Analysis. (n.d.). Retrieved March 19, 2020, from http://pantherdb.org/