This web page was produced as an assignment for Genetics 564, an undergraduate capstone course at UW-Madison.
What is Stargardt Disease?
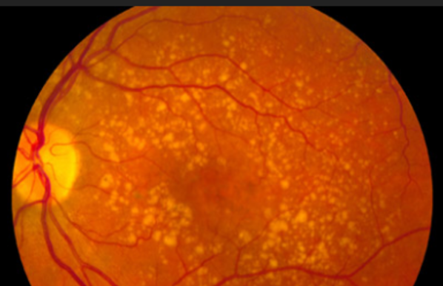
Stargardt disease is an autosomal recessive macular degeneration disease, affecting roughly 1 in 9,000 individuals and presents itself early in life. Macular degeneration is a type of illness that leads to loss of central vision progressively. In Stargardt disease this occurs with the buildup of lipofuscin globules in the eye, made up primarily of retinol and retinol derivative compounds. This buildup eventually leads to colorblindness and eventual complete vision loss.
What is ABCA4?
ABCA4 is the gene that is responsible for around 75% of cases of Stargardt disease. Located on chromosome one, this gene is made up of transmembrane regions, unknown regions, AAA ATPases, and ABC2 membrane 3s which themselves are made up of AAA ATPases and transmembrane regions.
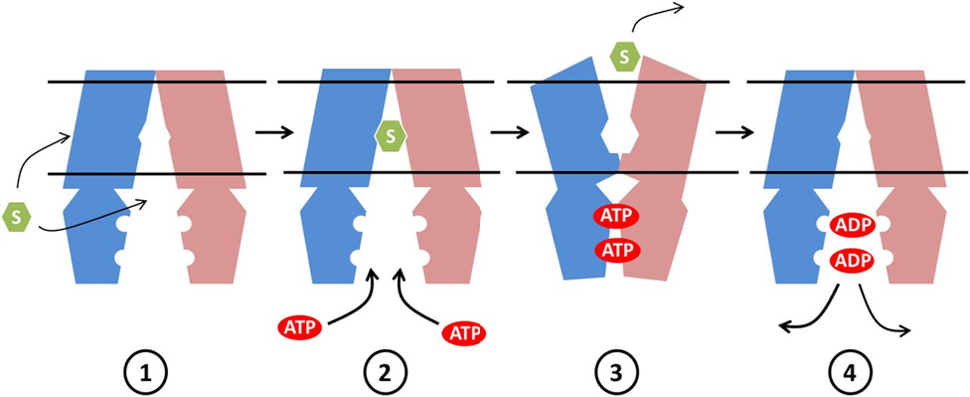
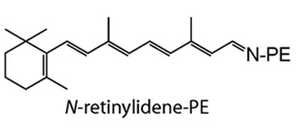
ABCA4 has a main molecular function as an ATP-binding cassette, taking in ATP in order to power the passage of a substrate across a cellular membrane. It's main cellular component is lipid transport across a membrane, of which it primarily transports N-retinylidene-PE. ABCA4 is associated mainly with the biological process of visual perception.
Homology of ABCA4
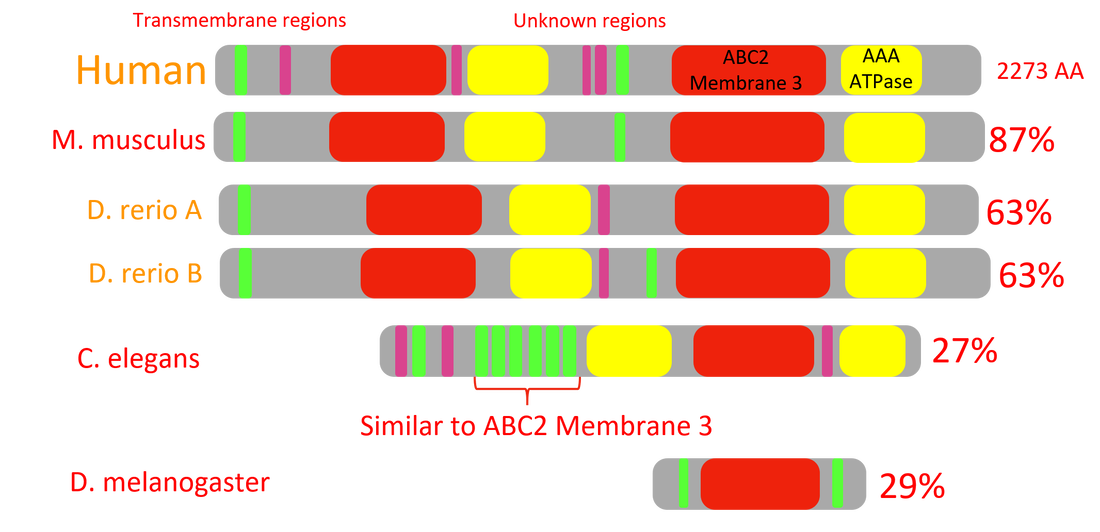
Figure 6: ABCA4 homology.
ABCA4 is highly conserved between species with similar eye types, like humans, mice, or zebrafish. It is less so conserved in species without eyes, like C. elegans, or with different eye types, like D. melanogaster.
ABCA4 protein interactions
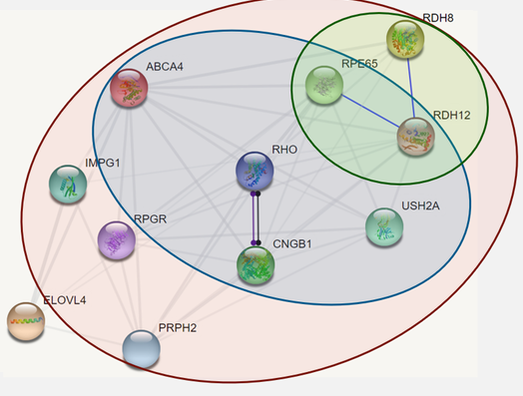
ABCA4 is highly connected with proteins associated with visual perception, retinal homeostasis, and retinol metabolism. What is most interesting is the lack of knowledge about modes of interaction between ABCA4 and retinol metabolism, considering that the ABCA4 nonfunctional Stargardt disease phenotype is primarily formed due to buildup of retinol in lipofuscin globules in the eye.
What is the gap in knowledge?
|
The role of ABCA4 in retinol metabolism is unclear. While there is an association between retinol metabolism proteins and ABCA4 the mode of interaction is unknown. This is surprising considering the main cause of ABCA4 responsible Stargardt disease phenotypes is the buildup of toxic retinol compounds in the eye.
|
Figure 8: Vitamin A1 (Retinol)
|
Why is zebrafish an excellent model species to study ABCA4?
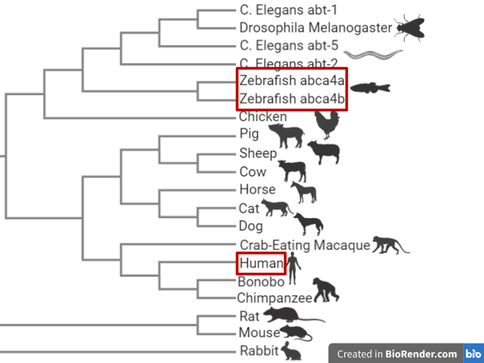
Figure 9: Phylogeny of ABCA4
|
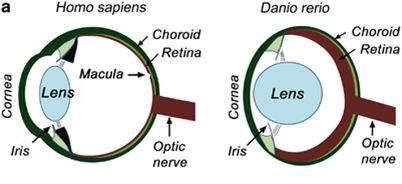
Despite the distance between zebrafish and human ABCA4 genes in phylogenetic terms, the ABCA4 gene is relatively well conserved between humans and zebrafish. Also the zebrafish and human eyes are very similar, allowing for closer similarity in phenotypes. Zebrafish also have a clear ABCA4 negative phenotype that can be assayed easily, as well as having rapid retinal development. Another model species that could have been used is the mouse, however the mouse does not have a clear ABCA4 phenotype and has a much less rapid retinal development.
|
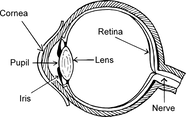
Figure 10: Human and zebrafish eyes.
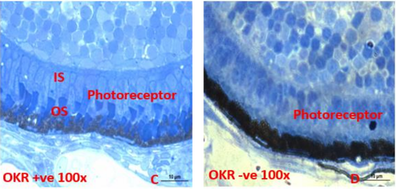
Figure 11: ABCA4 positive (left) and negative (right) phenotypes in the retina.
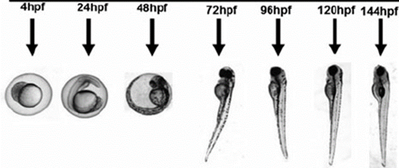
Figure 12: Zebrafish development.
|
What is the primary goal?
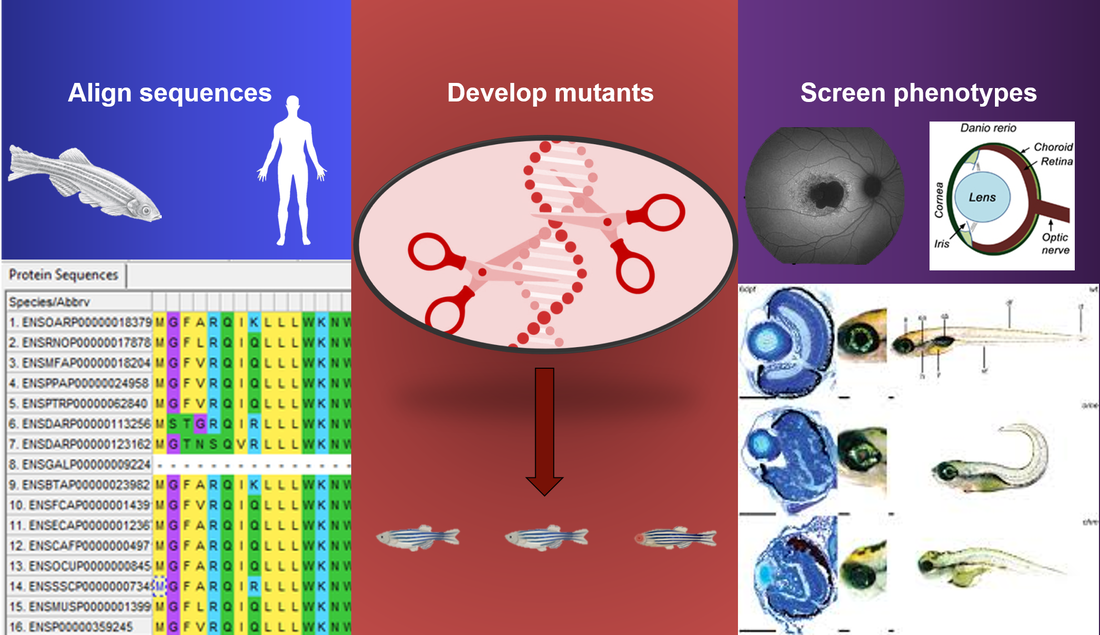
The primary goal is to determine the function of ABCA4 in retinol regulation during development. This will be done through analysis of domains important to the function of ABCA4, analysis of the chemical modulation of retinol metabolism, and by analyzing the expression of ABCA4 and retinol metabolism genes throughout development.
Aim 1: Determine domains important for retinol metabolism between human and Danio rerio ABCA4 genes.
Hypothesis: Danio rerio and human mutations in ABCA4 conserved regions will confer similar retinol metabolism phenotypes.
Approach: NCBI BLAST can be used to determine homologs between human ABCA4 and Danio rerio equivalents. This can be followed with Clustal Omega to identify conserved regions. The regions can then be disrupted with CRISPR-Cas9 and phenotypes can be analyzed and compared, in particular looking for abnormal accumulation of toxic retinoid compounds.
Rationale: Through analysis of phenotypes in Danio rerio ABCA4 conserved region mutations it can be determined how similar the mutant phenotypes are between human and Danio rerio, and which domains are critical for retinol metabolism. It can also orient us to differences we may have to account for between Danio rerio and humans.
Hypothesis: Danio rerio and human mutations in ABCA4 conserved regions will confer similar retinol metabolism phenotypes.
Approach: NCBI BLAST can be used to determine homologs between human ABCA4 and Danio rerio equivalents. This can be followed with Clustal Omega to identify conserved regions. The regions can then be disrupted with CRISPR-Cas9 and phenotypes can be analyzed and compared, in particular looking for abnormal accumulation of toxic retinoid compounds.
Rationale: Through analysis of phenotypes in Danio rerio ABCA4 conserved region mutations it can be determined how similar the mutant phenotypes are between human and Danio rerio, and which domains are critical for retinol metabolism. It can also orient us to differences we may have to account for between Danio rerio and humans.
Aim 2: Determine modulation of retinol metabolism.
Hypothesis: Retinol metabolism chemical pathways are disrupted in mutant phenotypes.
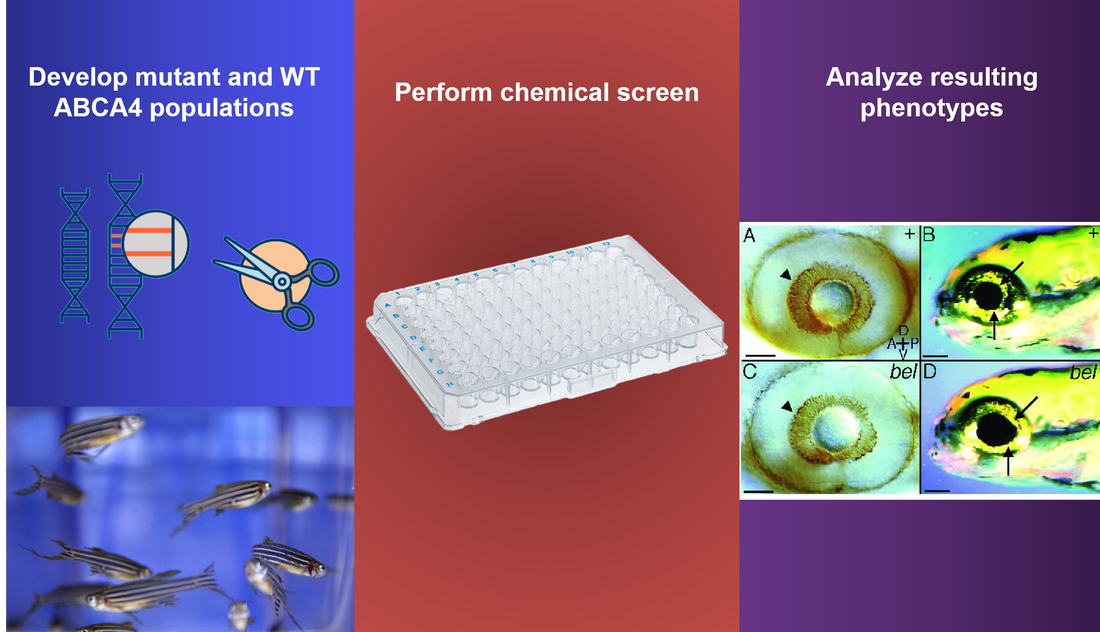
Approach: Nonfunctional ABCA4 and wild type Danio rerio populations can be produced, nonfunctional ABCA4 populations developed via CRISPR-Cas9 mutations, and put through a wide chemical screen with some of the chemicals chosen using the PubChem website. The following phenotypes can then be analyzed in the mutant and wild type Danio rerio subjects, looking for abnormal accumulation of toxic retinoid compounds in wild type populations as well as recovery in nonfunctional ABCA4 populations.
Rationale: Analysis of the resulting phenotypes, both wild type and mutant Danio rerio, can elucidate both chemicals that disrupt retinol metabolism pathways, as well as chemicals that are important for normal operation of retinol metabolism pathways.
Hypothesis: Retinol metabolism chemical pathways are disrupted in mutant phenotypes.
Approach: Nonfunctional ABCA4 and wild type Danio rerio populations can be produced, nonfunctional ABCA4 populations developed via CRISPR-Cas9 mutations, and put through a wide chemical screen with some of the chemicals chosen using the PubChem website. The following phenotypes can then be analyzed in the mutant and wild type Danio rerio subjects, looking for abnormal accumulation of toxic retinoid compounds in wild type populations as well as recovery in nonfunctional ABCA4 populations.
Rationale: Analysis of the resulting phenotypes, both wild type and mutant Danio rerio, can elucidate both chemicals that disrupt retinol metabolism pathways, as well as chemicals that are important for normal operation of retinol metabolism pathways.
Aim 3: Determine the expression of retinol metabolism genes through time.
Hypothesis: ABCA4 and other retinol metabolism genes change expression over the lifetime of species.
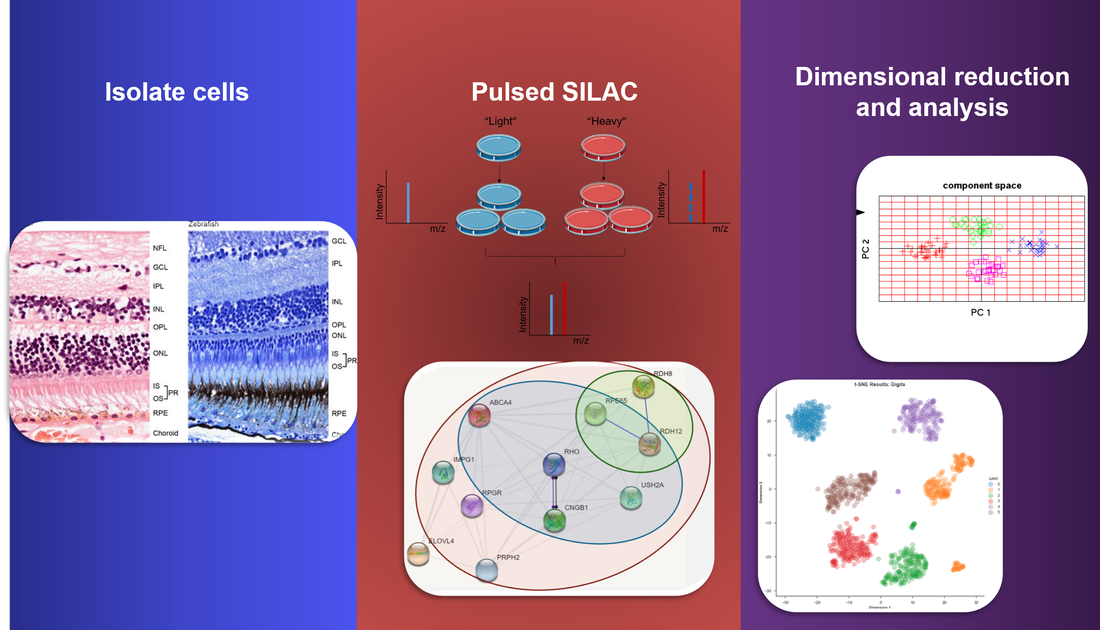
Approach: Retinal cells can be collected and cultured at different subject ages, from both nonfunctional ABCA4 and wild type populations, and subjected to SILAC with protein data collected. The data can then be normalized and analyzed through dimensionality reduction techniques such as PCA and t-SNE at each time point.
Rationale: Analysis of proteins could allow for an understanding of cell regulatory networks, response at different points in the life cycle, and protein-protein interactions. It would also allow for further understanding of ABCA4 in its regulatory network.
Hypothesis: ABCA4 and other retinol metabolism genes change expression over the lifetime of species.
Approach: Retinal cells can be collected and cultured at different subject ages, from both nonfunctional ABCA4 and wild type populations, and subjected to SILAC with protein data collected. The data can then be normalized and analyzed through dimensionality reduction techniques such as PCA and t-SNE at each time point.
Rationale: Analysis of proteins could allow for an understanding of cell regulatory networks, response at different points in the life cycle, and protein-protein interactions. It would also allow for further understanding of ABCA4 in its regulatory network.
Summary
Alignment of sequences and determination of mutant Zebrafish phenotypes may assist with understanding modes of retinol metabolism differing between species.
Performing chemical screens on wildtype and mutant Zebrafish can elucidate chemical modulation of retinol metabolism.
Understanding the expression of cells in the retina can further the understanding of ABCA4 interactions in retinol metabolism.
Performing chemical screens on wildtype and mutant Zebrafish can elucidate chemical modulation of retinol metabolism.
Understanding the expression of cells in the retina can further the understanding of ABCA4 interactions in retinol metabolism.
Future Directions
Some future directions past this study could include RNA or even single cell RNA sequencing to further study the interactions ABCA4 is involved with in the retina, and further analysis of chemicals that are found to be promising in recovering wild type phenotype from ABCA4 negative subjects could offer treatment options.
Presentation Drafts
| roughdraftpresentation2_25_20.pptx | |
| File Size: | 7696 kb |
| File Type: | pptx |
| roughdraftpresentation4_5_20.pdf | |
| File Size: | 2969 kb |
| File Type: | |
| roughdraftpresentation4_16_20.pdf | |
| File Size: | 3080 kb |
| File Type: | |
| finalpresentation4_25_20.pdf | |
| File Size: | 4337 kb |
| File Type: | |
Specific Aims Drafts
| timothybiewerheislerspecificaimsdraft4_15_20.docx | |
| File Size: | 20 kb |
| File Type: | docx |
| timothybiewerheislerspecificaimsdraft4_25_20.docx | |
| File Size: | 20 kb |
| File Type: | docx |